Do you have DNA matches that seem to belong in multiple genetic networks? Are you nervous that you’re seeing pedigree collapse or endogamy? Before you despair, realize that this phenomenon could be due to DNA matches sharing more than one common ancestral couple with each other, or what’s often called “multiple relationships.” In Diana’s post, Endogamy, Pedigree Collapse, and Multiple Relationships: What’s the Difference and Why Does it Matter, she talked briefly about multiple relationships and how it affects analysis of DNA matches. In this post I’ll share a more in-depth example of multiple relationships and discuss the effect on chromosome mapping, total amount of shared DNA, and network analysis.
Other Articles in this Series
Endogamy, Pedigree Collapse and Multiple relationships: What’s the Difference and Why does it matter? by Diana Elder, AG
How Multiple Relationships Affect DNA Match Analysis by Nicole Elder Dyer
Multiple Relationships in an African-American Case Study by Allison Kotter
The Effect of Pedigree Collapse on DNA Matching: A Case Study by Nicole Elder Dyer
Visualizing Complicated Relationships: Working with Pedigree Collapse, Multiple Relationships, and Endogamy by Steve Little
Strategies for Overcoming Endogamy by Nicole Elder Dyer
The Irish: Endogam-ish by Heidi Mathis
Endogamy: Ashkenazi Jewish Case by Heidi Mathis
The Richards and Robinson Families
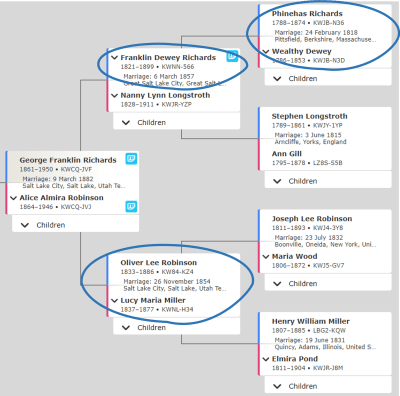
My example is from the Richards and Robinson families in my husband’s family tree. In the diagram below, you can see how Test Taker A (TTA) and Test Taker B (TTB) are double third cousins through both the Richards and Robinsons. One of their common ancestors, Franklin D. Richards, had two wives. TTA descends from Franklin D. Richards and Nanny Longstroth. TTB descends from Franklin D. Richards and a different wife, Rhoda Harriet Foss. Through the common ancestor Franklin D. Richards, TTA and TTB are half third cousins (H3C).
Franklin D. Richards’ son, George Franklin Richards, married Alice Almira Robinson. George’s half sister, Sarah Elizabeth Richards, married Alice’s brother, Loren Jay Robinson. So, TTA and TTB also share the common ancestral couple of Oliver Lee Robinson and Lucy Maria Miller. Through this ancestral couple, they are third cousins (3C).
Effect of Multiple Relationships on Chromosome Mapping
When mapping segments for matches with multiple relationships, it helps to map or “paint” as many matches as possible that do not have double or multiple relationships first. Once you have done that, you can overlay the double cousin segments to see if they overlap one of the ancestral couples you’ve already mapped. I’ll illustrate this with TTB, a match at MyHeritage. TTB shares 3 segments with TTA. When I overlay these shared segments on my chromosome map for TTA at DNA Painter, I see that two of them are overlapping Richards segments (gold) on chromosomes 5 and 7, and one is overlapping with a Robinson segment (hot pink) on chromosome 6.
 This is what I expected to see, knowing TTB is a double cousin sharing these two common ancestral couples. To paint these segments, I can copy and paste them one at a time into DNA Painter. The new segment from TTB on chromosome six is longer than the Robinson/Miller segment it overlaps. It could be helpful down the road to tentatively assign that segment to the Robinson/Miller group. In DNA Painter, you can change the certainty level and add notes for segments. Click “Add details about these match segments.” I used “possible” and then added a note about TTB being a double cousin. It’s also possible that there was a crossover at some point on that segment and TTB inherited a Richards segment adjacent to the Robinson segment, so my certainty level is lower.
This is what I expected to see, knowing TTB is a double cousin sharing these two common ancestral couples. To paint these segments, I can copy and paste them one at a time into DNA Painter. The new segment from TTB on chromosome six is longer than the Robinson/Miller segment it overlaps. It could be helpful down the road to tentatively assign that segment to the Robinson/Miller group. In DNA Painter, you can change the certainty level and add notes for segments. Click “Add details about these match segments.” I used “possible” and then added a note about TTB being a double cousin. It’s also possible that there was a crossover at some point on that segment and TTB inherited a Richards segment adjacent to the Robinson segment, so my certainty level is lower.
For the other two segments, I did the same except assigned them to the Richards/Dewey group (parents of Franklin D. Richards). When you change the certainty level to likely or possible, the color of the painted segment becomes a little bit lighter or faded.
Non-Overlapping Segments
I found another descendant of Loren Robinson and Sarah Elizabeth Richards in the MyHeritage database, who I’ll call Test Taker C (TTC). He shares four segments with TTA. When I overlaid these segments, two did not overlap with known segments. I needed a way to label these double cousin segments. Reviewing the tree of TTA, I thought at first that I could add them to the Richards/Robinson group. However, that would indicate that it included all ancestors on George Franklin Richards and Alice Almira Robinson’s lines – but there is one line they don’t represent – Nanny Longstroth. It would be more accurate to call the unknown segments “Richards/Dewey or Robinson/Miller,” since Franklin Dewey Richards is the MRCA on the Richards side.
I went ahead and labelled the segment group “Richards/Dewey or Robinson/Miller,” and gave it a reddish pink color. On chromosome 2, the segment shared with TTC covered an area that hadn’t been painted at all yet. Although I can’t be sure if the segment is Richards/Dewey or Robinson/Miller, at least I have it narrowed down to those two options and painted more of the chromosome! (Isn’t it exciting to see the red message at the top of DNA Painter go up to a higher percentage of the genome painted?) Although the adjacent segments are Robinson/Brown and Robinson/Miller, that doesn’t necessarily mean the unknown segment is also Robinson. It’s best to keep both options in the label until we learn more.
The takeaway from this exercise is that mapping segments for genetic cousins with an unknown relationship can reveal multiple relationships. If they overlap two separate ancestral couples from different lines, you may need to investigate that possibility. Build our their tree on all lines to find additional MRCAs.
Effect of Multiple Relationships on Amount of Shared DNA
Now that we saw the effect of multiple relationships on segments, let’s review how it affects the total amount of shared DNA. In Diana’s post about endogamy, pedigree collapse, and multiple relationships, she talked about how usually people who have multiple relationships don’t share so much more DNA that they are bumped into a different relationship category. Let’s test that out with the Richards family.
TTA and TTB share 103.9 cM.
TTA and TTC share 88.1 cM.
Both of those amounts are within the range of 3rd cousin. The mean is 73 cM and the standard deviation is 43 cM. One standard deviation above the mean is 116 cM. Although these cousins are both third cousins and half third cousins, they don’t share so much extra DNA that it changes them into the 2nd cousin range.
For the half 3rd cousin relationship, the amounts are definitely on the high end. The mean for Half 3C is 48 cM and the standard deviation is 33 cM. One standard deviation above the mean would be 81 cM, so 88 and 103 cM both fall above that. If all we knew is that they were half third cousins, and didn’t realize they shared another 3rd cousin relationship, these numbers might tip us off that we should look for another relationship.
Effect of Multiple Relationships on Network Analysis
Now let’s review the effect of multiple relationships on genetic networks. The MyHeritage AutoCluster report for TTA shows some overlapping between the first two clusters. The first cluster, red, is primarily Richards/Dewey descendants. The second cluster, orange, includes Robinson/Miller descendants, as well as the descendants of Loren Robinson and Sarah Richards – the double cousins TTB and TTC – and several additional double cousins. This causes many gray cells which are not in a cluster, which indicate shared matches between the red and orange clusters. When you see gray cells, it indicates those two clusters are connected somehow. In this case, they are connected because of multiple relationships between TTA and several of the matches in the orange and red clusters, and not that the orange cluster is a further back ancestral couple along the same line as the red cluster.
Network Graph of AncestryDNA Matches
Test Taker A has even more Richards and Robinson matches at AncestryDNA. Many descendants of George Franklin Richards and Alice Robinson have tested at AncestryDNA. Many descendants of Loren Robinson and Sarah Richards have also tested at AncestryDNA. The network graph below shows the effect of all these test takers having multiple relationships with each other. There is such a strong connection between the Richards/Dewey Cluster and Robinson/Miller cluster that they are almost completely stuck together. Often you won’t be able to distinguish clusters when many people in those clusters share multiple relationships with each other.
The takeaway? When analyzing a network graph or cluster chart, and some clusters are very closely connected, check for multiple relationships. Although it may appear that there are too many connections to figure out which clusters are relevant, don’t worry. The challenge of multiple relationships between matches is not as difficult as dealing with pedigree collapse or endogamy. You can do it!
Dive in and figure out which matches are connected to both clusters, and find the MRCAs. Once you know which matches have multiple relationships, you can label them as such and focus on the matches that don’t have that issue. If you have segment data for those matches, try chromosome mapping with DNA painter to help you hypothesize which segment came from which MRCA couple. Good luck!
Learn More
Diahan Southard offers an online course about dealing with endogamy. Diahan has a wonderful way of teaching about DNA that helps you gain confidence! Learn more and register here: Start Untangling Your Family Tree | Endogamy & DNA Course
To learn how to create network graphs like the ones in this article, join our online course, Research Like a Pro with DNA. In this course, we take you step by step through organizing and analyzing your matches and applying DNA evidence to focused research questions.



















Leave a Reply
Thanks for the note!